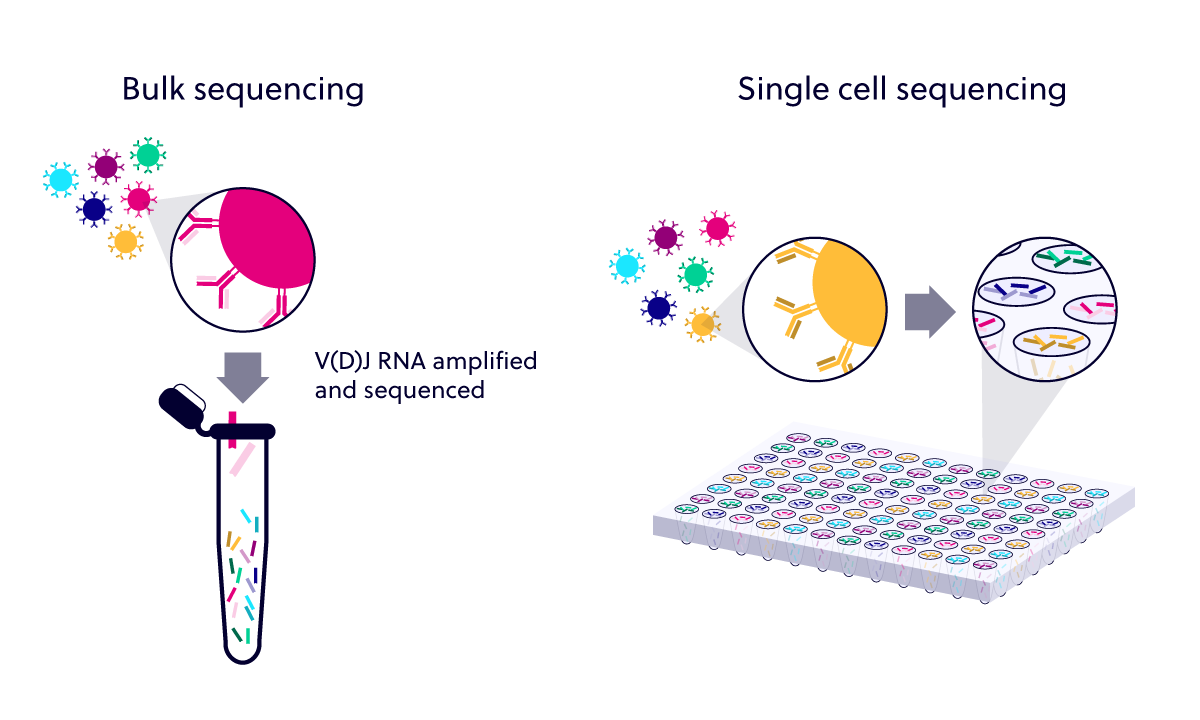
scFlow combines best-practice approaches for processing sc/snRNA-seq datasets see ‘Materials and methods’ for a detailed explanation of these steps. This pipeline included the removal of empty droplets, nuclei with low read counts and doublets, followed by embedding and integration of cells from separate samples and cell typing. To gain advantage of these recent approaches, we used scFlow ( Khozoie et al., 2021) to reprocess the authors’ data. As the field has matured since the authors’ work was published, dataset integration has become a common step in sc-RNA-seq protocols and is recommended by some to remove confounding sources of variation ( Heumos et al., 2023 Amezquita et al., 2020 Tran et al., 2020). Moreover, the authors did not integrate the cells from different individuals to account for batch effects. This is indicative of low cell quality ( Ilicic et al., 2016) however, we believe the authors’ QC approach may not capture all of these low-quality cells. Firstly, in relation to their processing approach, the authors discussed the high percentages of mitochondrial reads and low number of reads per cell present in their data. Our questions of Mathys et al., 2019 focus around their data processing and their differential expression (DE) analysis ( Figure 1).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
